 I’m currently taking a data mining class, and I’m trying to do as much of the coding in the class using R for practice. We are using Weka 3 which so far is proving to be a pretty neat (FREE) data mining tool. To meet my goal of improving my R skills while taking this class I needed to find a way to read .arff data files from Weka into R so that I can access the data to run the assigned tasks.
I’m currently taking a data mining class, and I’m trying to do as much of the coding in the class using R for practice. We are using Weka 3 which so far is proving to be a pretty neat (FREE) data mining tool. To meet my goal of improving my R skills while taking this class I needed to find a way to read .arff data files from Weka into R so that I can access the data to run the assigned tasks.
The first option, which is what most of my classmates are doing, is of course is to load the data into Weka and convert it to a CSV file and then load that file into R using the read.csv() command. I was curious to see if I could do it more directly.
After some research I found that there are several options, one of which is the RWeka library. When I tried to install the RWeka package into RStudio (on Mac OS X) I ran into a few issues and errors mostly involving Java. Here is the solution I found after a lovely Sunday morning jumping from thread to the thread on the ever helpful Stack Overflow:
1. Relink R with the installed Java Library on OS X.
Open the terminal and run this command:
sudo R CMD javareconf
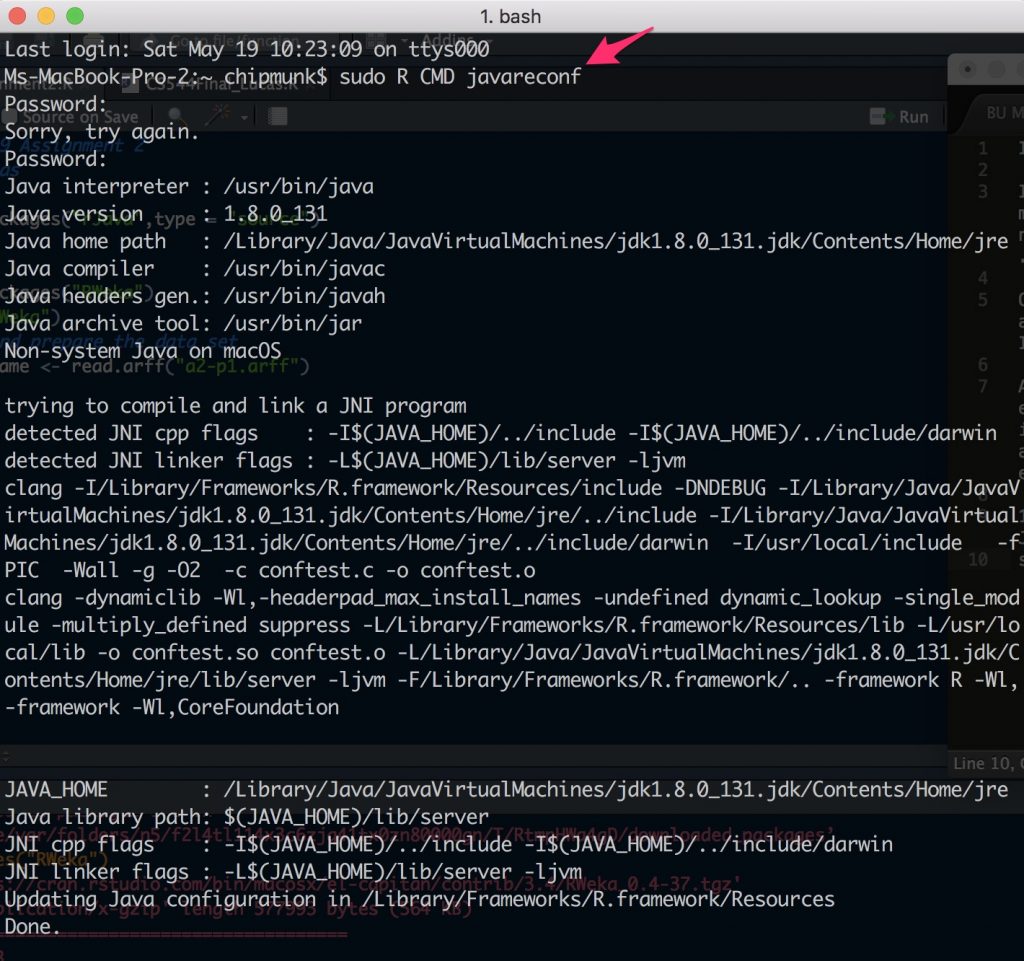
2. Install the rJava package in R.
I’m using RStudio here, but either way the command would be:
install.packages(“rJava”,type = “source”)
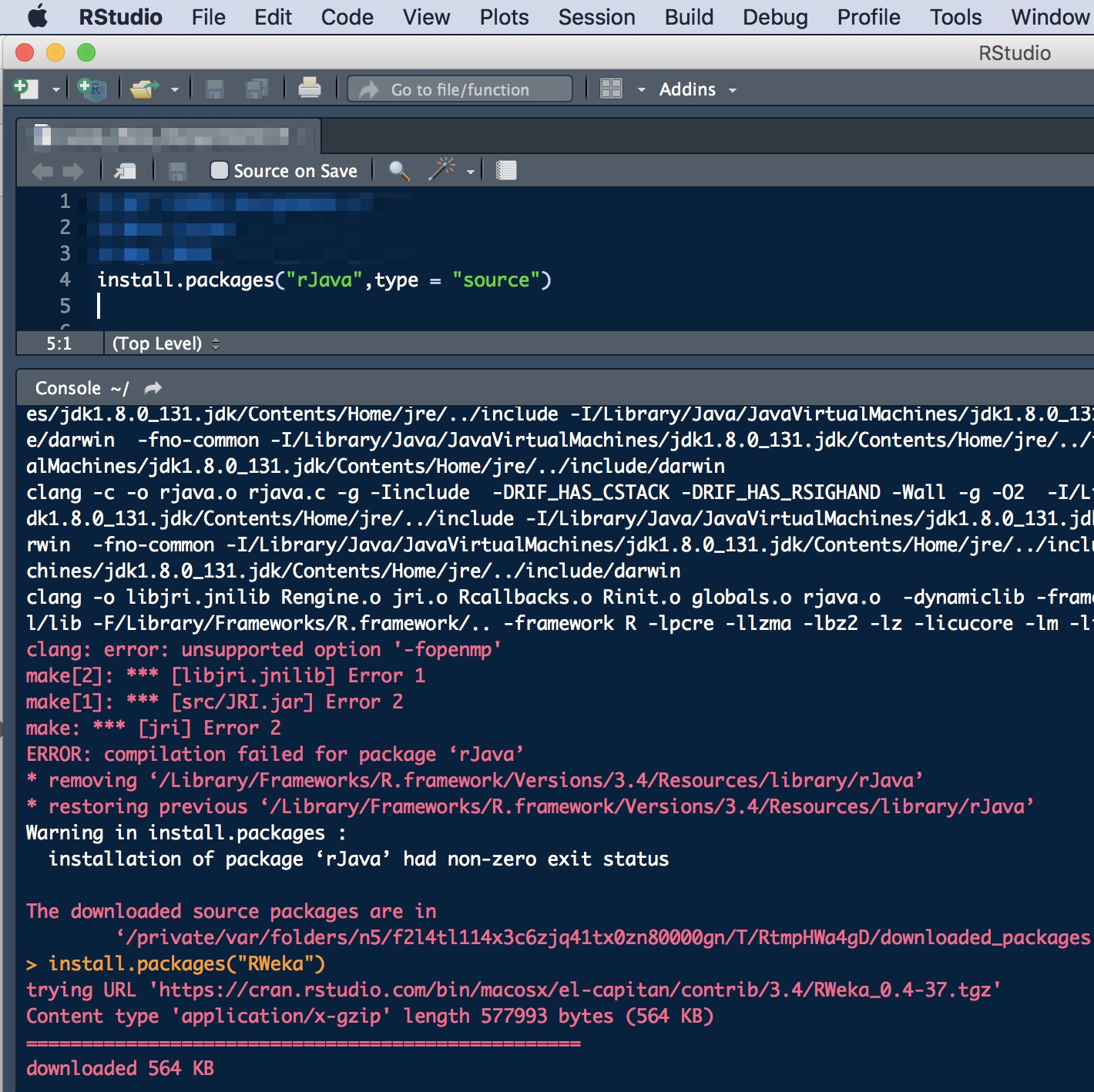
I was a bit worried about all those errors but then I realized I had already installed rJava before, hence the “restorng previous…” message. The “non-zero exit status” was probably a result of the previous errors, and I could not figure out what if there was another possible reason for it, so I decided to keep going.
3. Install the RWeka package in R (or RStudio):
The last step was to install the RWeka package, and then load the library. I have put those commands together here, but I ran the install first, got the success message, then loaded the library.
install.packages(“RWeka”)
library(“RWeka”)
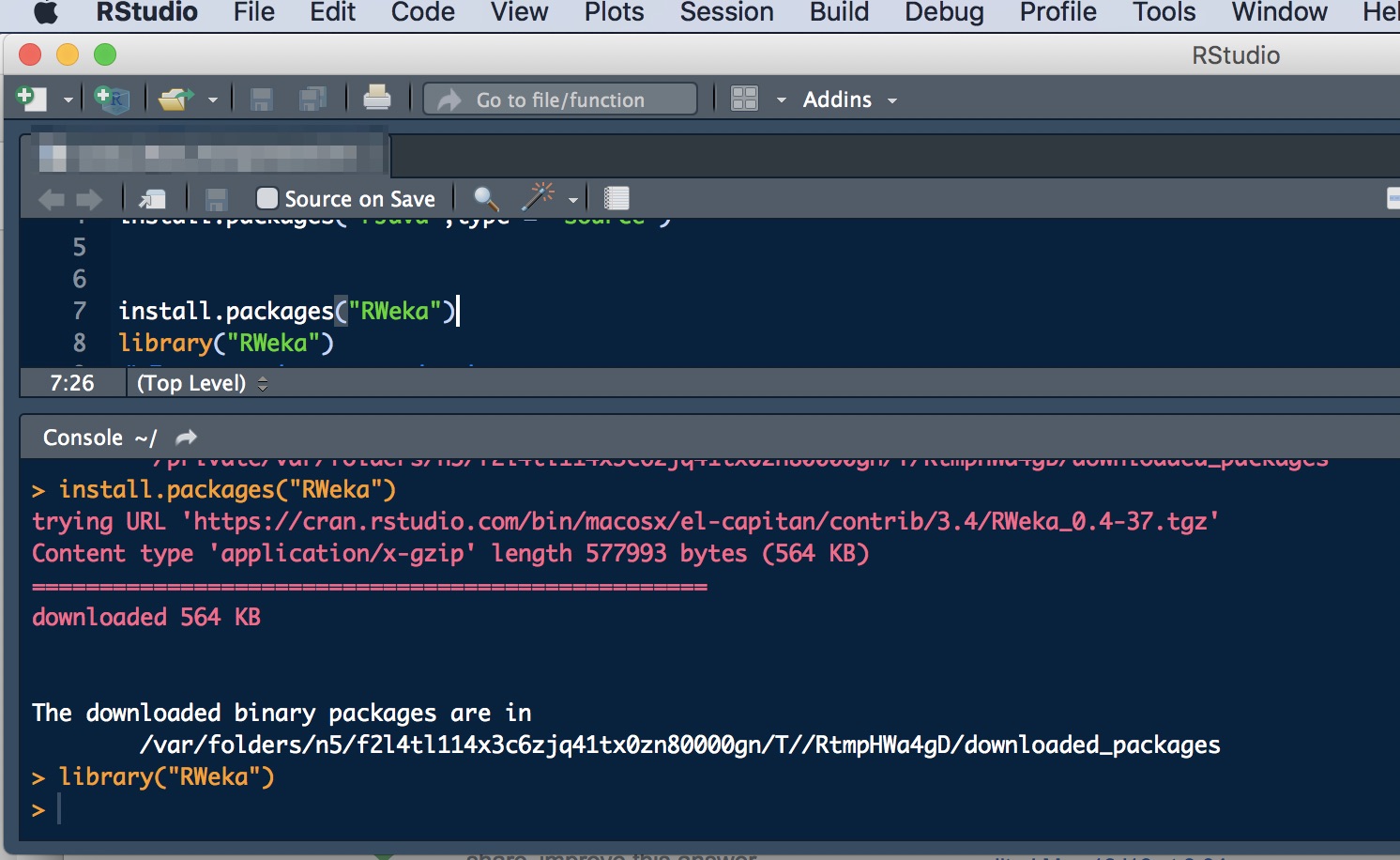
After that I was able to load the arff data file using the read.arff command and everything is working perfectly!
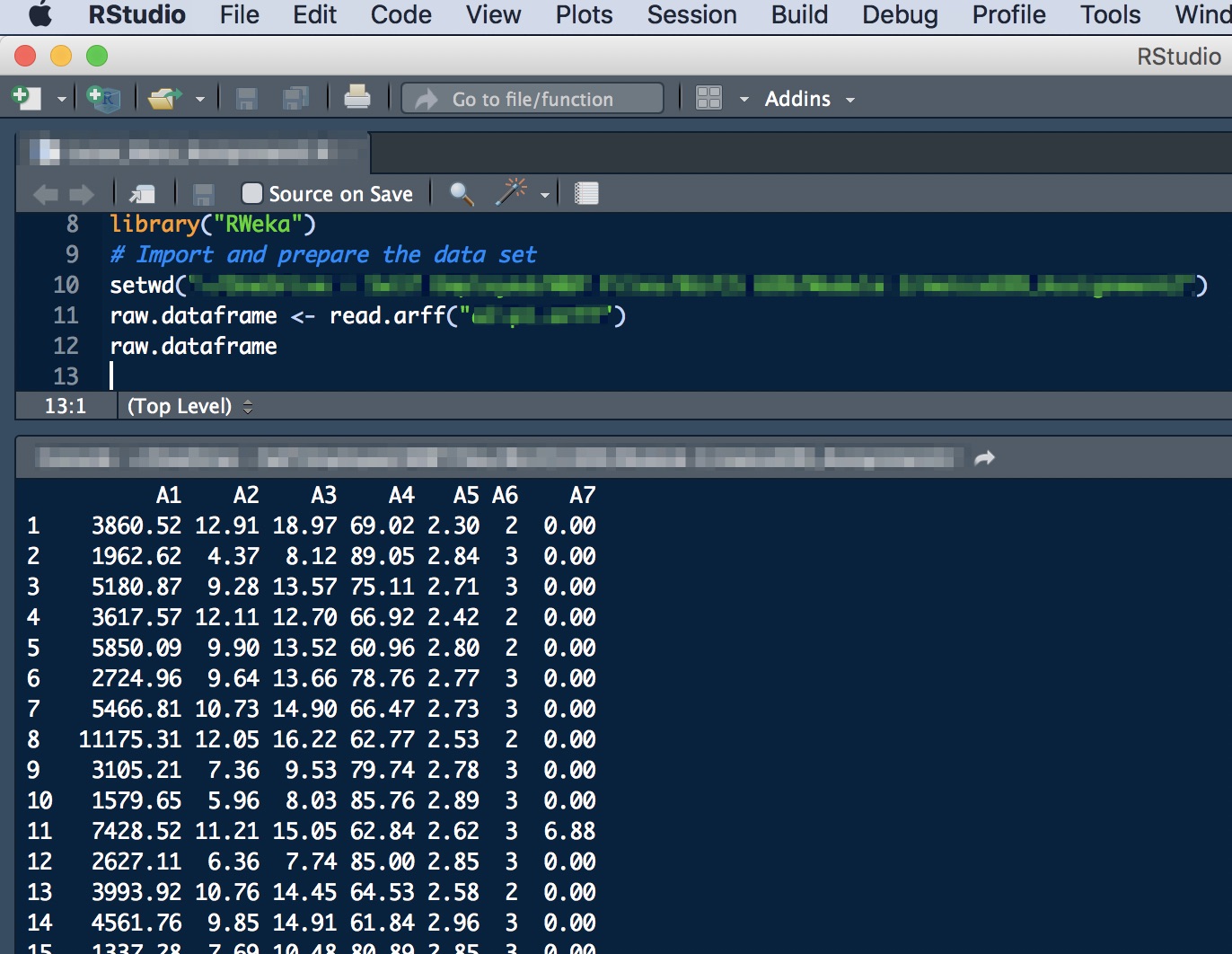
Hope this is helpful!
References:
https://stackoverflow.com/questions/35179151/cannot-load-r-xlsx-package-on-mac-os-10-11
https://stackoverflow.com/questions/30738974/rjava-load-error-in-rstudio-r-after-upgrading-to-osx-yosemite
Thank you for your write. Entering the terminal command resulted in a long series of error messages:
missing xcrun at: /Library/Developer/CommandLineTools/usr/bin/xcrun
sh: –version: command not found
using SDK: ‘NA’
xcrun: error: invalid active developer path (/Library/Developer/CommandLineTools), missing xcrun at: /Library/Developer/CommandLineTools/usr/bin/xcrun
Warning messages:
1: In system(paste(MAKE, p1(paste(“-f”, shQuote(makefiles))), “compilers”), :
running command ‘make -f ‘Makevars’ -f ‘/Library/Frameworks/R.framework/Resources/etc/Makeconf’ -f ‘/Library/Frameworks/R.framework/Resources/share/make/shlib.mk’ compilers’ had status 1
2: In system2(“xcrun”, “–show-sdk-path”, TRUE, TRUE) :
running command ”xcrun’ –show-sdk-path 2>&1′ had status 1
Unable to compile a JNI program
I will keep looking for an answer.